Finding the Right Medication Codes for Research
Assembling a comprehensive list of medications used to treat a specific disease is harder than it should be. There’s no single place to look it up. The ATC (Anatomical Therapeutic Chemical) classification system organizes drugs by their pharmacological mechanism — but that tells a researcher what a drug is, not what it treats. MED-RT (Medication Reference Terminology), the NLM’s successor to the older NDF-RT, maintains curated “may treat” relationships between drugs and conditions, but coverage is uneven and the vocabulary is coarse.
For a well-understood condition like hypertension, this might feel manageable — ACE inhibitors, ARBs, calcium channel blockers, beta-blockers, and diuretics are the obvious starting points. But even then, that’s five different ATC therapeutic groups to traverse. For something like rheumatoid arthritis, where the treatment landscape spans conventional DMARDs, biologics targeting different cytokines, JAK inhibitors, corticosteroids, and NSAIDs, the picture gets complicated fast. And for any condition where off-label use is common or where the standard of care has shifted recently, manual assembly is a recipe for gaps.
We needed something better for our work at Outcomes Insights. So we built it.
The Traditional Approach and Its Limitations
The standard pharmacological classification systems are organized around what a drug is, rather than what it does for a patient. ATC classifies drugs into a hierarchy based on the organ system they act on and their pharmacological mechanism. That’s useful for understanding drug classes, but it doesn’t directly answer the question a researcher typically starts with “what medications are used to treat this disease?”
MED-RT gets closer. It maintains explicit “may treat” relationships between drugs and conditions. But these relationships are curated at the ingredient level, coverage varies across therapeutic areas, and combination products are largely absent — of the roughly 3,800 combination ingredients in RxNorm, almost none have “may treat” links. If a patient is prescribed amlodipine/benazepril for hypertension, MED-RT knows about amlodipine and benazepril individually, but it doesn’t know about the combination product that actually appears on the claim.
The deeper problem is structural. Classification systems organize medications by chemistry and mechanism. Researchers and analysts need to find medications by therapeutic intent. These are related perspectives, but they aren’t the same thing — and the gap between them is where medications get missed.
Starting from What Drugs Actually Say They Do
We took a different approach. Instead of working backward from classification systems, we started with what drugs say they do — literally. Every FDA-approved medication comes with a Structured Product Label (SPL) that includes an “Indications and Usage” section. This is the authoritative statement of what the drug is approved to treat, in clinical language.
We built a pipeline that extracts structured therapeutic relationships from these indication statements across roughly 2,000 ingredients with available labels. Each extraction captures a condition-action-outcome triple:
- What condition does this drug address?
- What does it do?
- What’s the intended outcome?
An LLM reads the raw label text and produces these structured extractions, which are then validated against the source.
Here’s the key step: we encode each extracted indication as a vector embedding using BioLORD-2023, a biomedical language model. This transforms clinical statements like “treatment of primary hypercholesterolemia” and “reduction of elevated total cholesterol” into mathematical representations that capture their clinical meaning. Those embeddings go into a vector database — one entry per indication sentence, linked back to the originating ingredient.
This creates something that didn’t exist before: a searchable index of what medications actually do, expressed in the language of clinical practice rather than the language of pharmacological classification.
Searching by Disease, Not by Drug Class
With this index in place, a disease query — “type 2 diabetes,” “rheumatoid arthritis,” “treatment-resistant depression” — returns a ranked list of every ingredient whose approved indications are semantically close to the query.
The system uses the same class of embedding technology behind our ICD-10 Code Set Builder, but applied to a different corpus. Instead of encoding ICD-10 code descriptions, we’re encoding drug indication statements. The search doesn’t require exact keyword matches. A query for “diabetic medications for obesity” surfaces GLP-1 receptor agonists like semaglutide and liraglutide — drugs classified under ATC code A10BJ (“Blood Glucose Lowering Drugs / GLP-1 analogues”) whose labels describe weight management indications. It also picks up tirzepatide from a different ATC subclass (A10BX). These are diabetes drugs by classification, but the system ranks them by relevance to obesity because their indication text matches the query semantically, not lexically (i.e., as a text match). (Note that ingredients that are in the class, but have no indication information that is semantically relevant, are listed as “class only” and do not have any indication score.)
Results come back grouped by ATC therapeutic class, so the familiar pharmacological organization is preserved as context. The figure below shows this in action — the GLP-1 class appears with semaglutide, liraglutide, and others ranked by both class membership and indication relevance, with the system auto-selecting the most relevant ingredients.
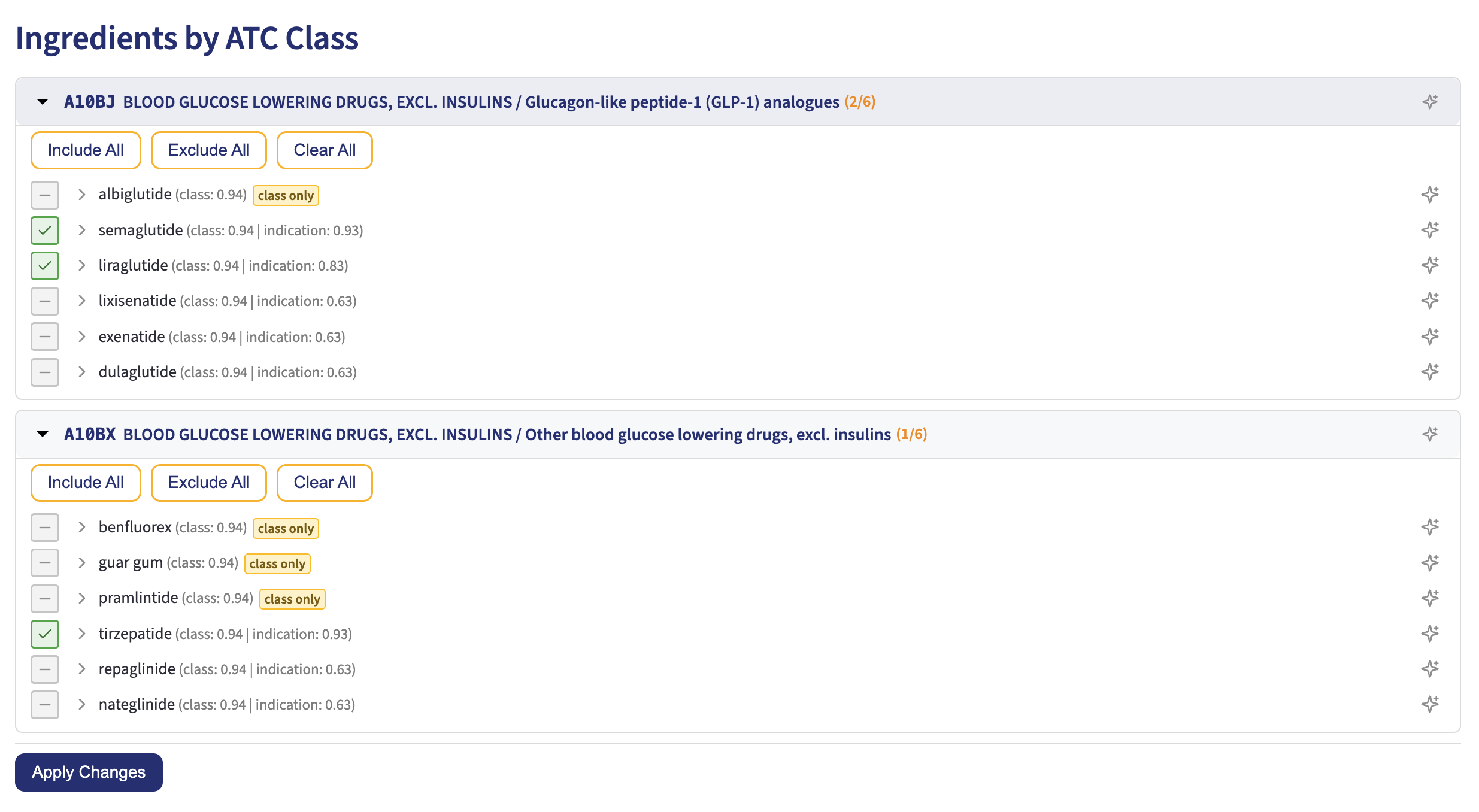
But the tool also surfaces drugs from other ATC groups when their indications match the query — a statin with a specific diabetes-related indication, for example, or an antihypertensive with outcome data in diabetic patients.
This dual organization — searched by disease, displayed by drug class — bridges the gap between how researchers think about medications and how pharmacology organizes them.
Two Modes of Search
Not every question starts with a disease. Sometimes a researcher already knows the drug class of interest and needs to enumerate its members. The tool supports both modes:
Condition-based search encodes the disease query as an embedding and searches the indication corpus. This is the mode that captures therapeutic intent — it finds drugs based on what they treat, regardless of where they sit in the ATC hierarchy.
Class-based search encodes a pharmacological class name — “SGLT2 inhibitors,” “TNF-alpha inhibitors,” “SSRIs” — and searches a separate collection of ATC class embeddings. This is useful when the user already has a therapeutic class in mind and wants to enumerate its members quickly.
In practice, many queries blend both perspectives. A search for “GLP-1 agonists for obesity” triggers a class-focused search that identifies the GLP-1 receptor agonist class, then cross-references indication embeddings to rank members by relevance to obesity specifically — distinguishing between GLP-1 drugs approved for weight management and those approved only for diabetes.
From Ingredients to Billable Codes
Finding the right ingredients is only half the problem. The practical deliverable for most use cases is a set of billable codes — NDC codes for pharmacy claims and HCPCS J-codes for physician-administered drugs. The system follows each ingredient through the RxNorm hierarchy to its dispensable products and their associated NDC codes, then cross-references the CMS NDC-to-HCPCS crosswalk for injectable and infused medications.
Each ingredient in the results shows its NDC and HCPCS counts, available product forms, and representative indication text. Users can include or exclude individual ingredients, and the system pre-selects likely relevant drugs using a combination of semantic similarity scoring and an optional AI pre-check that reviews the selections in clinical context.
Why This Matters
Medication identification is foundational to a broad range of healthcare research — pharmacoepidemiology, outcomes studies, formulary analysis, health economics.
The traditional approach — browsing ATC hierarchies and curated drug-disease databases — works, but it organizes information around the drug rather than around the patient’s condition. Our approach inverts the starting point: begin with what you want to treat, and let the system find the medications that treat it. The pharmacological classification is still there, providing structure and context, but it’s no longer the only way in.
This is the second application we’ve built on this foundation. The first, our Code Set Builder for ICD-10-CM, uses the same semantic search principles to identify diagnosis codes. Both tools share a conviction that clinical intent (i.e., what am I looking for?) should be the starting point for code set construction, with classification hierarchies serving as guardrails rather than the primary navigation mechanism.
 by Outcomes Insights, Inc.
by Outcomes Insights, Inc.